Multiple Correspondence Analysis with Orthogonal Instrumental Variables
MCAoiv.RdMultiple Correspondence Analysis with Orthogonal Instrumental Variables
MCAoiv(X, Z, excl = NULL, row.w = NULL, ncp = 5)Arguments
- X
data frame with only factors
- Z
data frame of instrumental variables, which can be numeric or factors. It must have the same number of rows as
X.- excl
numeric vector indicating the indexes of the "junk" categories (default is NULL). See
getindexcator useijunkinteractive function to identify these indexes. It may also be a character vector of junk categories, specified in the form "namevariable.namecategory" (for instance "gender.male").- row.w
Numeric vector of row weights. If NULL (default), a vector of 1 for uniform row weights is used.
- ncp
number of dimensions kept in the results (by default 5)
Details
Multiple Correspondence Analysis with Orthogonal Instrumental Variables consists in three steps :
1. Specific MCA of Y, keeping all the dimensions of the space
2. Computation of one linear regression for each dimension in the specific MCA, with individual coordinates as response and all variables in X as explanatory variables.
3. Principal Component Analysis of the set of residuals from the regressions in 2.
Note
If there are NAs in Y, these NAs will be automatically considered as junk categories. If one desires more flexibility, Y should be recoded to add explicit factor levels for NAs and then excl option may be used to select the junk categories.
Value
An object of class PCA from FactoMineR package, with X as supplementary variables, and an additional item :
- ratio
the share of inertia not explained by the instrumental variables
.
References
Bry X., 1996, Analyses factorielles multiples, Economica.
Lebart L., Morineau A. et Warwick K., 1984, Multivariate Descriptive Statistical Analysis, John Wiley and sons, New-York.)
Examples
library(FactoMineR)
data(tea)
mcaoiv <- MCAoiv(tea[,1:18], tea[,19:22])
mcaoiv$ratio
#> [1] 0.9429018
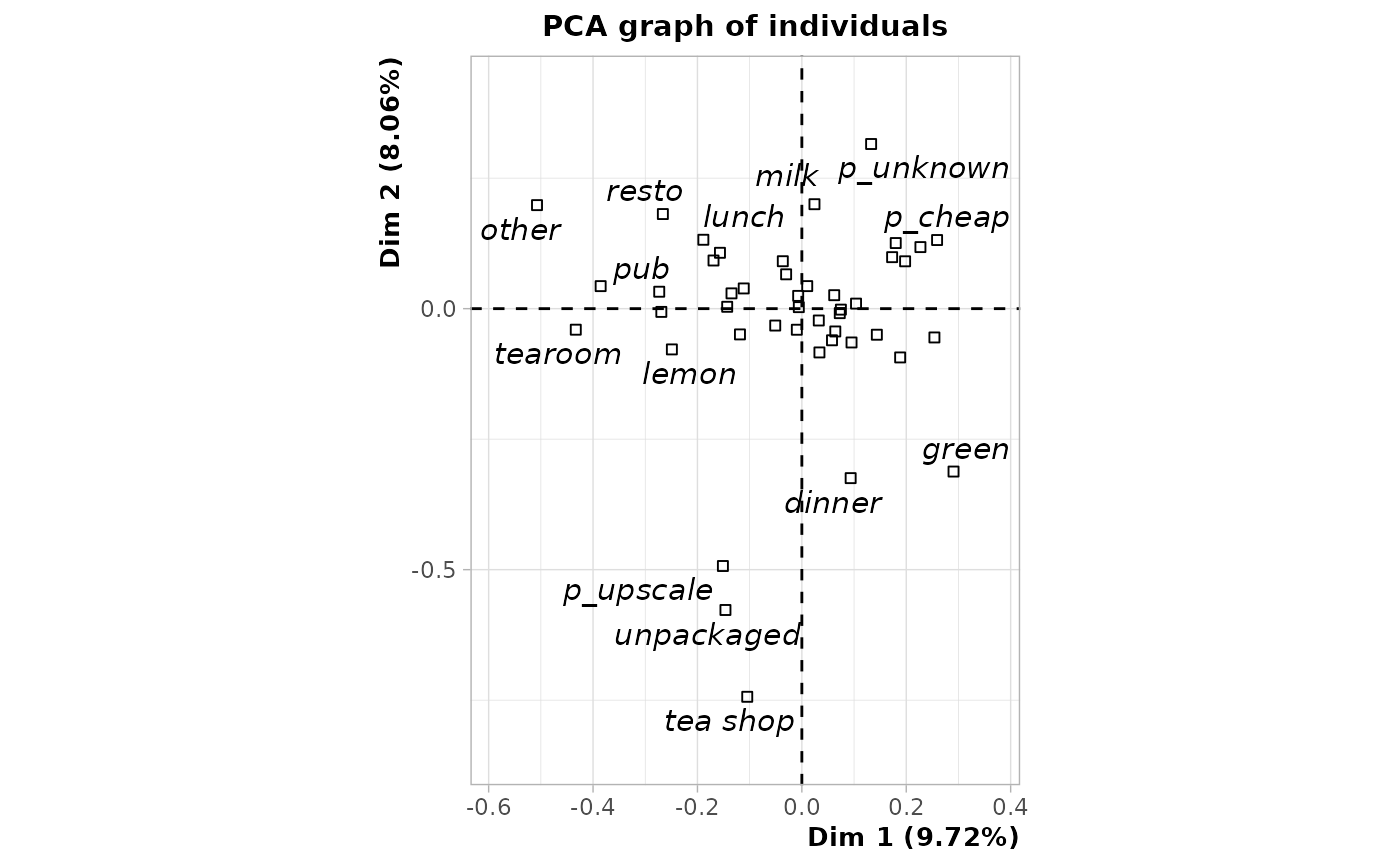
plot(mcaoiv, choix = "ind", invisible = "ind", col.quali = "black")
#> Warning: ggrepel: 18 unlabeled data points (too many overlaps). Consider increasing max.overlaps